Note
Go to the end to download the full example code.
Profiling your models¶
Do you feel like your model is too slow? Do you want to make it faster? Instead of guessing which part of the code is responsible for any slowdown, you should profile your code to learn how much time is spent in each function and where to focus any optimization efforts.
In this tutorial you’ll learn how to profile your model using PyTorch profiler, how to read the output of the profiler, and how to add your own labels for new functions/steps in your model forward function.
from typing import Dict, List, Optional
import ase.build
import matplotlib.pyplot as plt
import numpy as np
import torch
from metatensor.torch import Labels, TensorBlock, TensorMap
from metatensor.torch.atomistic import (
MetatensorAtomisticModel,
ModelCapabilities,
ModelMetadata,
ModelOutput,
System,
)
from metatensor.torch.atomistic.ase_calculator import MetatensorCalculator
When profiling your code, it is important to run the model on a representative system to ensure you are actually exercising the behavior of your model at the right scale. Here we’ll use a relatively large system with many atoms.
primitive = ase.build.bulk(name="C", crystalstructure="diamond", a=3.567)
atoms = ase.build.make_supercell(primitive, 10 * np.eye(3))
print(f"We have {len(atoms)} atoms in our system")
We have 2000 atoms in our system
We will use the same HarmonicModel as in the previous tutorial as our machine learning potential.
Click to see the definition of HarmonicModel
class HarmonicModel(torch.nn.Module):
def __init__(self, force_constant: float, equilibrium_positions: torch.Tensor):
"""Create an ``HarmonicModel``.
:param force_constant: force constant, in ``energy unit / (length unit)^2``
:param equilibrium_positions: torch tensor with shape ``n x 3``, containing the
equilibrium positions of all atoms
"""
super().__init__()
assert force_constant > 0
self.force_constant = force_constant
self.equilibrium_positions = equilibrium_positions
def forward(
self,
systems: List[System],
outputs: Dict[str, ModelOutput],
selected_atoms: Optional[Labels],
) -> Dict[str, TensorMap]:
# if the model user did not request an energy calculation, we have nothing to do
if "energy" not in outputs:
return {}
# we don't want to worry about selected_atoms yet
if selected_atoms is not None:
raise NotImplementedError("selected_atoms is not implemented")
if outputs["energy"].per_atom:
raise NotImplementedError("per atom energy is not implemented")
# compute the energy for each system by adding together the energy for each atom
energy = torch.zeros((len(systems), 1), dtype=systems[0].positions.dtype)
for i, system in enumerate(systems):
assert len(system) == self.equilibrium_positions.shape[0]
r0 = self.equilibrium_positions
energy[i] += torch.sum(self.force_constant * (system.positions - r0) ** 2)
# add metadata to the output
block = TensorBlock(
values=energy,
samples=Labels("system", torch.arange(len(systems)).reshape(-1, 1)),
components=[],
properties=Labels("energy", torch.tensor([[0]])),
)
return {
"energy": TensorMap(keys=Labels("_", torch.tensor([[0]])), blocks=[block])
}
model = HarmonicModel(
force_constant=3.14159265358979323846,
equilibrium_positions=torch.tensor(atoms.positions),
)
capabilities = ModelCapabilities(
outputs={
"energy": ModelOutput(quantity="energy", unit="eV", per_atom=False),
},
atomic_types=[6],
interaction_range=0.0,
length_unit="Angstrom",
supported_devices=["cpu"],
dtype="float32",
)
metadata = ModelMetadata()
wrapper = MetatensorAtomisticModel(model.eval(), metadata, capabilities)
wrapper.export("exported-model.pt")
/home/runner/work/metatensor/metatensor/python/examples/atomistic/3-profiling.py:126: DeprecationWarning: `export()` is deprecated, use `save()` instead
wrapper.export("exported-model.pt")
If you are trying to profile your own model, you can start here and create a
MetatensorCalculator with your own model.
atoms.calc = MetatensorCalculator("exported-model.pt")
Before trying to profile the code, it is a good idea to run it a couple of times to allow torch to warmup internally.
atoms.get_forces()
atoms.get_potential_energy()
3.770593615115558e-09
Profiling energy calculation¶
Now we can run code using torch.profiler.profile() to collect statistic on
how long each function takes to run. We randomize the positions to force ASE to
recompute the energy of the system
atoms.positions += np.random.rand(*atoms.positions.shape)
with torch.profiler.profile() as energy_profiler:
atoms.get_potential_energy()
print(energy_profiler.key_averages().table(sort_by="self_cpu_time_total", row_limit=10))
-------------------------------------------------- ------------ ------------ ------------ ------------ ------------ ------------
Name Self CPU % Self CPU CPU total % CPU total CPU time avg # of Calls
-------------------------------------------------- ------------ ------------ ------------ ------------ ------------ ------------
Model::forward 44.26% 393.000us 57.21% 508.000us 508.000us 1
ASECalculator::prepare_inputs 21.17% 188.000us 24.66% 219.000us 219.000us 1
ASECalculator::convert_outputs 9.01% 80.000us 10.36% 92.000us 46.000us 2
ASECalculator::compute_neighbors 3.15% 28.000us 3.15% 28.000us 28.000us 1
aten::_to_copy 2.93% 26.000us 5.29% 47.000us 5.222us 9
aten::to 1.91% 17.000us 6.42% 57.000us 3.562us 16
aten::copy_ 1.80% 16.000us 1.80% 16.000us 1.778us 9
aten::mul 1.58% 14.000us 1.58% 14.000us 14.000us 1
aten::arange 1.58% 14.000us 2.70% 24.000us 12.000us 2
aten::sum 1.46% 13.000us 1.46% 13.000us 13.000us 1
-------------------------------------------------- ------------ ------------ ------------ ------------ ------------ ------------
Self CPU time total: 888.000us
There are a couple of interesting things to see here. First the total runtime of the
code is shown in the bottom; and then the most costly functions are visible on top,
one line per function. For each function, Self CPU refers to the time spent in
this function excluding any called functions; and CPU total refers to the time
spent in this function, including called functions.
For more options to record operations and display the output, please refer to the official documentation for PyTorch profiler.
Here, Model::forward indicates the time taken by your model’s forward().
Anything starting with aten:: comes from operations on torch tensors, typically
with the same function name as the corresponding torch functions (e.g.
aten::arange is torch.arange()). We can also see some internal functions
from metatensor, with the name staring with MetatensorAtomisticModel:: for
MetatensorAtomisticModel; and ASECalculator:: for
ase_calculator.MetatensorCalculator.
If you want to see more details on the internal steps taken by your model, you can add
torch.profiler.record_function()
(https://pytorch.org/docs/stable/generated/torch.autograd.profiler.record_function.html)
inside your model code to give names to different steps in the calculation. This is
how we are internally adding names such as Model::forward or
ASECalculator::prepare_inputs above.
Profiling forces calculation¶
Let’s now do the same, but computing the forces for this system. This mean we should
now see some time spent in the backward() function, on top of everything else.
atoms.positions += np.random.rand(*atoms.positions.shape)
with torch.profiler.profile() as forces_profiler:
atoms.get_forces()
print(forces_profiler.key_averages().table(sort_by="self_cpu_time_total", row_limit=10))
------------------------------------------------------- ------------ ------------ ------------ ------------ ------------ ------------
Name Self CPU % Self CPU CPU total % CPU total CPU time avg # of Calls
------------------------------------------------------- ------------ ------------ ------------ ------------ ------------ ------------
torch::jit::(anonymous namespace)::DifferentiableGra... 36.22% 732.000us 43.69% 883.000us 883.000us 1
Model::forward 18.65% 377.000us 26.72% 540.000us 540.000us 1
ASECalculator::prepare_inputs 8.66% 175.000us 9.70% 196.000us 196.000us 1
ASECalculator::run_backward 5.59% 113.000us 54.53% 1.102ms 1.102ms 1
aten::mul 5.24% 106.000us 5.54% 112.000us 28.000us 4
ASECalculator::convert_outputs 5.20% 105.000us 6.19% 125.000us 62.500us 2
aten::copy_ 2.82% 57.000us 2.82% 57.000us 3.800us 15
<backward op> 1.93% 39.000us 7.47% 151.000us 151.000us 1
aten::pow 1.68% 34.000us 2.38% 48.000us 24.000us 2
aten::_to_copy 1.48% 30.000us 3.86% 78.000us 6.500us 12
------------------------------------------------------- ------------ ------------ ------------ ------------ ------------ ------------
Self CPU time total: 2.021ms
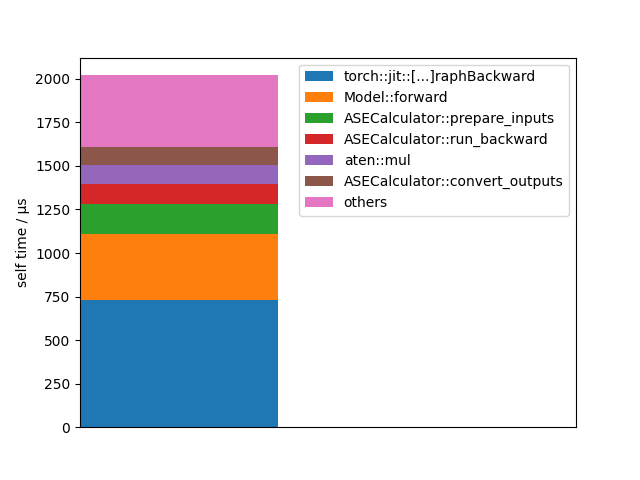
Let’s visualize this data in an other way:
events = forces_profiler.key_averages()
events = sorted(events, key=lambda u: u.self_cpu_time_total, reverse=True)
total_cpu_time = sum(map(lambda u: u.self_cpu_time_total, events))
bottom = 0.0
for event in events:
self_time = event.self_cpu_time_total
name = event.key
if len(name) > 30:
name = name[:12] + "[...]" + name[-12:]
if self_time > 0.03 * total_cpu_time:
plt.bar(0, self_time, bottom=bottom, label=name)
bottom += self_time
else:
plt.bar(0, total_cpu_time - bottom, bottom=bottom, label="others")
break
plt.legend()
plt.xticks([])
plt.xlim(0, 1)
plt.ylabel("self time / µs")
plt.show()
Total running time of the script: (0 minutes 0.260 seconds)