Note
Go to the end to download the full example code.
Batched simulations with TorchSim¶
TorchSim supports batching multiple systems into a single SimState
for efficient parallel evaluation on GPU.
MetatomicModel handles this
transparently.
Setup¶
We reuse the same minimal model from Getting started with TorchSim. The model must produce differentiable energy so that forces/stress can be computed via autograd.
from typing import Dict, List, Optional
import ase.build
import matplotlib.pyplot as plt
import torch
import torch_sim as ts
from metatensor.torch import Labels, TensorBlock, TensorMap
import metatomic.torch as mta
from metatomic_torchsim import MetatomicModel
class HarmonicEnergy(torch.nn.Module):
"""Harmonic restraint: E = k * sum(positions^2)."""
def __init__(self, k: float = 0.1):
super().__init__()
self.k = k
def forward(
self,
systems: List[mta.System],
outputs: Dict[str, mta.ModelOutput],
selected_atoms: Optional[Labels] = None,
) -> Dict[str, TensorMap]:
energies: List[torch.Tensor] = []
for system in systems:
e = self.k * torch.sum(system.positions**2)
energies.append(e.reshape(1, 1))
energy = torch.cat(energies, dim=0)
block = TensorBlock(
values=energy,
samples=Labels("system", torch.arange(len(systems)).reshape(-1, 1)),
components=[],
properties=Labels("energy", torch.tensor([[0]])),
)
return {
"energy": TensorMap(keys=Labels("_", torch.tensor([[0]])), blocks=[block])
}
capabilities = mta.ModelCapabilities(
length_unit="Angstrom",
atomic_types=[13, 29], # Al, Cu
interaction_range=0.0,
outputs={"energy": mta.ModelOutput(unit="eV")},
supported_devices=["cpu"],
dtype="float64",
)
atomistic_model = mta.AtomisticModel(
HarmonicEnergy(0.1).eval(), mta.ModelMetadata(), capabilities
)
model = MetatomicModel(atomistic_model, device="cpu")
Creating a batched state¶
Pass a list of ASE Atoms objects to initialize_state:
atoms_list = [
ase.build.bulk("Cu", "fcc", a=3.6, cubic=True),
ase.build.bulk("Cu", "fcc", a=3.65, cubic=True),
ase.build.bulk("Al", "fcc", a=4.05, cubic=True),
]
sim_state = ts.initialize_state(atoms_list, device=model.device, dtype=model.dtype)
print("Total atoms in batch:", sim_state.n_atoms)
Total atoms in batch: 12
Evaluating the batch¶
A single forward call evaluates all systems:
results = model(sim_state)
print("Energy shape:", results["energy"].shape) # [n_systems]
print("Forces shape:", results["forces"].shape) # [n_total_atoms, 3]
print("Stress shape:", results["stress"].shape) # [n_systems, 3, 3]
Energy shape: torch.Size([3])
Forces shape: torch.Size([12, 3])
Stress shape: torch.Size([3, 3, 3])
The output shapes reflect the batch:
results["energy"]has shape[n_systems]– one energy per systemresults["forces"]has shape[n_total_atoms, 3]– all atoms concatenatedresults["stress"]has shape[n_systems, 3, 3]– one 3x3 tensor per system
print("Per-system energies:", results["energy"])
Per-system energies: tensor([1.9440, 1.9984, 2.4604], dtype=torch.float64)
How system_idx works¶
SimState tracks which atom belongs to which system via the
system_idx tensor. For three 4-atom systems, system_idx looks
like:
print("system_idx:", sim_state.system_idx)
system_idx: tensor([0, 0, 0, 0, 1, 1, 1, 1, 2, 2, 2, 2])
MetatomicModel.forward uses this to split the batched positions and
types into per-system System objects before calling the underlying
model.
Batch consistency¶
Energies computed in a batch match those computed individually. This is guaranteed because each system gets its own neighbor list and independent evaluation:
individual_energies = []
for atoms in atoms_list:
state = ts.initialize_state(atoms, device=model.device, dtype=model.dtype)
res = model(state)
individual_energies.append(res["energy"].item())
print("Batched: ", [e.item() for e in results["energy"]])
print("Individual:", individual_energies)
plt.scatter(individual_energies, results["energy"].cpu().numpy())
plt.plot(
[min(individual_energies), max(individual_energies)],
[min(individual_energies), max(individual_energies)],
"k--",
)
plt.xlabel("Individual energies")
plt.ylabel("Batched energies")
plt.show()
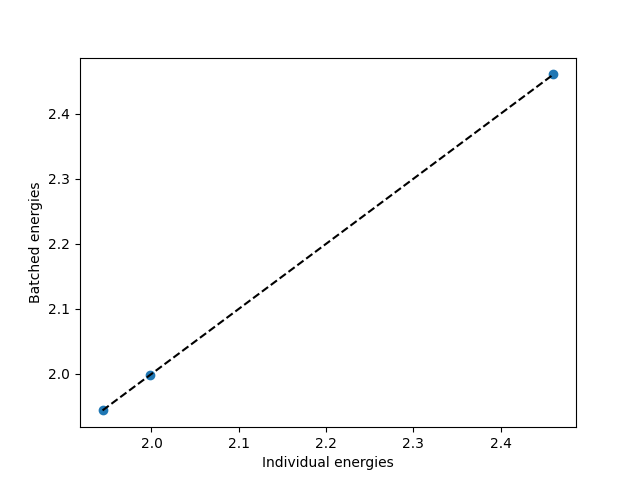
Batched: [1.9440000000000002, 1.9983750000000002, 2.460375]
Individual: [1.9440000000000002, 1.9983750000000002, 2.460375]
Performance considerations¶
Batching is most beneficial on GPU, where the neighbor list computation and model forward pass can run in parallel across systems. On CPU, the speedup comes from reduced Python overhead (one call instead of N).
For very large systems or many small ones, adjust the batch size to fit in GPU memory. TorchSim does not impose a maximum batch size, but each system gets its own neighbor list, so memory scales with the sum of per-system sizes.
Total running time of the script: (0 minutes 0.062 seconds)